 
So without actually doing anything to the source code (though it’s still nice to have to understand it) you can, say, loop through the different thresholding methods and see which one works for your project. The knime-cellprofiler integration communicates with CellProfiler using knime-bridge, handing over the images as ImgPlus objects. Which is awesome, since my compiled version works, and since (related breakthrough) you can edit the pipeline (.cp) files in a text editor (or, say, programmatically in Python). But! The small, trivial breakthrough is just that you can just run the compiled version in command line mode too: CellProfiler.exe -c -r -i ~/my_image_directory -o ~/my_output_directory -p ~/my_pipe.mat Per-object features: A CellProfiler pipeline can perform one or more segmentations which partition the image into many objects. python-loky: Robust and reusable executor for joblib. Which is all well and good except, despite my downloading and installing all the dozen or so libraries and things needed for CellProfiler, I still can’t actually get it to compile. fieldbioinformatics: pipeline for working with virus sequencing data sequenced with nanopore. To run CellProfiler 2.0 without the graphical user interface, use this: python CellProfiler.py -c -r -i ~/my_image_directory -o ~/my_output_directory -p ~/my_pipe.mat There are instructions on the CellProfiler Wiki for how to run CP from the command line:įor a full list of command-line options, run CellProfiler like this: python CellProfiler.py -help I noticed that many of the posts and questions around the CP-KNIME integration are a few years old and so was wondering whether the latest version(s) of CP still support(s) this integration? I’m runningĬellProfiler Integration 0.3.3.Brilliant! This small, trivial breakthrough has just saved me a ton of time. New and improved algorithms: for neuron image analysis, CellProfiler 2.0 includes operations to enhance neurites and to measure their branching, and algorithms for neuron-specific metrics are in development. 
The ‘Path to CP Installation’ is set correctly in KNIME preferences.Īny help with troubleshooting would be much appreciated! The same error occurs regardless of which sample workflow I use. This is helpful when you have a very large dataset. Or not so large scale analyses that you prefer to automate One of the benefits of running CellProfiler in batch is that you can split your analysis. 322, Pipeline Pilot, SVDetect, eFindSitePPI, PIQA, biojs-model. Getting Started with CellProfiler in Batch Running CellProfiler in batch mode is the ideal way to automate large scale analyses. WARN KNIPLogService TypeError: unicode not allowed, use setsockopt_string 166, Agp2amos, BSDB, CoNCoS, KNIME, Electron density server at Uppsala university EDS. WARN KNIPLogService File "zmq\backend\cython\socket.pyx", line 413, in .t The ‘Path to CP Installation’ is set correctly in KNIME preferences. WARN KNIPLogService File "cellprofiler\knime_bridge.py", line 117, in run WARN KNIPLogService File "threading.py", line 932, in _bootstrap_inner Z-stacks, time series, image masks), but it’s important to input them carefully so that CellProfiler can parse them correctly. To see the contents of a meta node (grey nodes) you can Double-Click on it, which opens a new tab with the nodes contained in this meta node. You may have to scroll down to see all contents of this tutorial. General Information Press the green double arrow above to run the complete workflow. CellProfiler can load many different data types (e.g. Run CellProfiler Colocalization Example in KNIME. WARN KNIPLogService Traceback (most recent call last): These modules are crucial for any CellProfiler pipeline because they define how images are loaded and organized in CellProfiler for downstream analysis. I’m trying the sample workflows and keep getting the following error when running the ‘CellProfiler Pipeline Executor’ node: WARN KNIPLogService Exception in thread Knime bridge server: KNIME Image Processing - CellProfiler Integration. The relevant code can be found in these two repositories: GitHub knime-ip/knip-cellprofiler. Export your data to a spreadsheet or database. The knime-cellprofiler integration communicates with CellProfiler using knime-bridge, handing over the images as ImgPlus objects. Process images automatically even millions. Adjust the settings to measure the phenotypes of interest in your images. Finally, it measures the number of overlapping spots per nucleus. 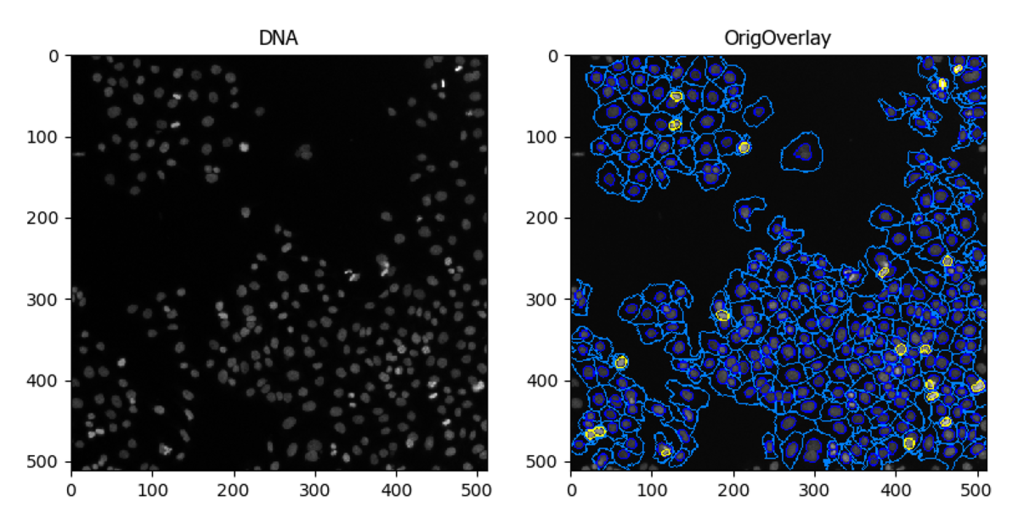
It then assigns each spot to a nucleus and counts the number per nucleus. I am trying to get the CellProfiler integration in KNIME to work. Designed for biologists Load an example CellProfiler pipeline, a series of image-processing modules. For some more background on my setup: I have a Cellprofiler pipeline that detects nuclei, red spots and green spots.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
